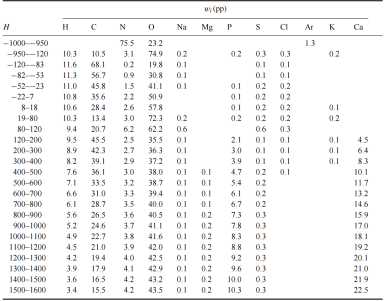
This represents the initial approach in Geant4, where patient materials are defined rather than roughly approximated as only 10 materials, and density variations can be controlled using densitydiff. In any case, the current set of 40 materials is a reasonably achievable solution for me at this stage, and it feels more realistic than approaches based purely on ICRU or NIST materials.
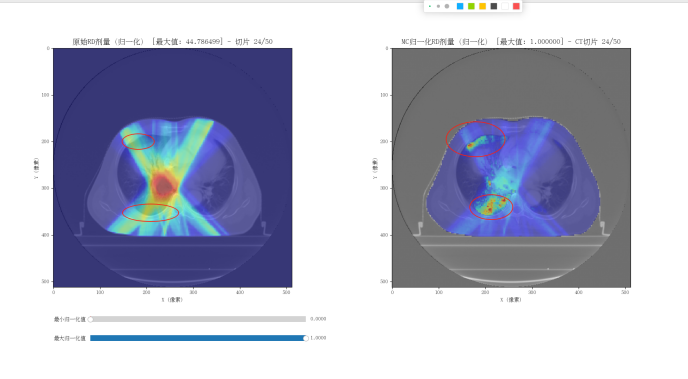
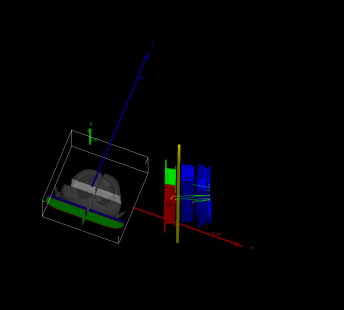
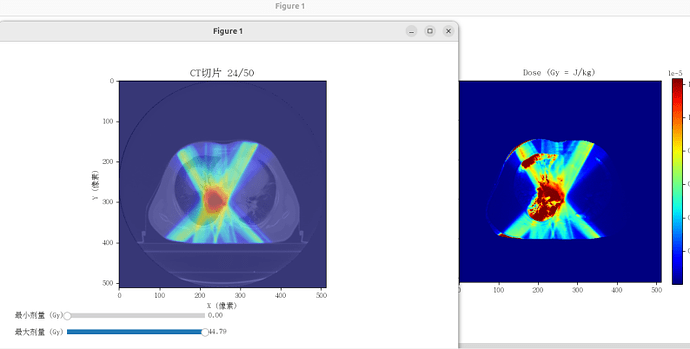
My goal is first to obtain a Monte Carlo dose distribution for an IMRT plan. With sufficient reference data available, I can then further refine the model, since my research is not intended for a commercial product. The current simulation will likely take about 10 more days to complete.G4Element* elC = new G4Element(name = “Carbon”, symbol = “C”, z = 6.0, a = 12.011 * g / mole);
G4Element* elH = new G4Element(name = “Hydrogen”, symbol = “H”, z = 1.0, a = 1.008 * g / mole);
G4Element* elN = new G4Element(name = “Nitrogen”, symbol = “N”, z = 7.0, a = 14.007 * g / mole);
G4Element* elO = new G4Element(name = “Oxygen”, symbol = “O”, z = 8.0, a = 16.00 * g / mole);
G4Element* elNa =
new G4Element(name = “Sodium”, symbol = “Na”, z = 11.0, a = 22.98977 * g / mole);
G4Element* elMg =
new G4Element(name = “Magnesium”, symbol = “Mg”, z = 12.0, a = 24.3050 * g / mole);
G4Element* elP =
new G4Element(name = “Phosphorus”, symbol = “P”, z = 15.0, a = 30.973976 * g / mole);
G4Element* elS = new G4Element(name = “Sulfur”, symbol = “S”, z = 16.0, a = 32.065 * g / mole);
G4Element* elCl =
new G4Element(name = “Chlorine”, symbol = “Cl”, z = 17.0, a = 35.453 * g / mole);
G4Element* elK =
new G4Element(name = “Potassium”, symbol = “K”, z = 19.0, a = 30.0983 * g / mole);
G4Element* elFe = new G4Element(name = “Iron”, symbol = “Fe”, z = 26, a = 56.845 * g / mole);
G4Element* elCa = new G4Element(name = “Calcium”, symbol = “Ca”, z = 20.0, a = 40.078 * g / mole);
G4Element* elZn = new G4Element(name = “Zinc”, symbol = “Zn”, z = 30.0, a = 65.382 * g / mole);
// 需要先定义 12 个元素,包括 Ar(例子代码里没建 Ar,要补上)
G4Element* elAr = new G4Element(name=“Argon”,symbol=“Ar”, z=18.0,a=39.948 * g/mole);
// Creating Materials :
G4int numberofElements;
// Air
fAir = new G4Material(“Air”, 1.290 * mg / cm3, numberofElements = 2);
fAir->AddElement(elN, 0.7);
fAir->AddElement(elO, 0.3);
// Soft tissue (ICRP - NIST)
G4Material* softTissue = new G4Material(“SoftTissue”, 1.00 * g / cm3, numberofElements = 13);
softTissue->AddElement(elH, 10.4472 * perCent);
softTissue->AddElement(elC, 23.219 * perCent);
softTissue->AddElement(elN, 2.488 * perCent);
softTissue->AddElement(elO, 63.0238 * perCent);
softTissue->AddElement(elNa, 0.113 * perCent);
softTissue->AddElement(elMg, 0.0113 * perCent);
softTissue->AddElement(elP, 0.113 * perCent);
softTissue->AddElement(elS, 0.199 * perCent);
softTissue->AddElement(elCl, 0.134 * perCent);
softTissue->AddElement(elK, 0.199 * perCent);
softTissue->AddElement(elCa, 0.023 * perCent);
softTissue->AddElement(elFe, 0.005 * perCent);
softTissue->AddElement(elZn, 0.003 * perCent);
// Lung Inhale
G4Material* lunginhale =
new G4Material(“LungInhale”, density = 0.217 * g / cm3, numberofElements = 9);
lunginhale->AddElement(elH, 0.103);
lunginhale->AddElement(elC, 0.105);
lunginhale->AddElement(elN, 0.031);
lunginhale->AddElement(elO, 0.749);
lunginhale->AddElement(elNa, 0.002);
lunginhale->AddElement(elP, 0.002);
lunginhale->AddElement(elS, 0.003);
lunginhale->AddElement(elCl, 0.002);
lunginhale->AddElement(elK, 0.003);
// Lung exhale
G4Material* lungexhale =
new G4Material(“LungExhale”, density = 0.508 * g / cm3, numberofElements = 9);
lungexhale->AddElement(elH, 0.103);
lungexhale->AddElement(elC, 0.105);
lungexhale->AddElement(elN, 0.031);
lungexhale->AddElement(elO, 0.749);
lungexhale->AddElement(elNa, 0.002);
lungexhale->AddElement(elP, 0.002);
lungexhale->AddElement(elS, 0.003);
lungexhale->AddElement(elCl, 0.002);
lungexhale->AddElement(elK, 0.003);
// Adipose tissue
G4Material* adiposeTissue =
new G4Material(“AdiposeTissue”, density = 0.967 * g / cm3, numberofElements = 7);
adiposeTissue->AddElement(elH, 0.114);
adiposeTissue->AddElement(elC, 0.598);
adiposeTissue->AddElement(elN, 0.007);
adiposeTissue->AddElement(elO, 0.278);
adiposeTissue->AddElement(elNa, 0.001);
adiposeTissue->AddElement(elS, 0.001);
adiposeTissue->AddElement(elCl, 0.001);
// Brain (ICRP - NIST)
G4Material* brainTissue = new G4Material(“BrainTissue”, 1.03 * g / cm3, numberofElements = 13);
brainTissue->AddElement(elH, 11.0667 * perCent);
brainTissue->AddElement(elC, 12.542 * perCent);
brainTissue->AddElement(elN, 1.328 * perCent);
brainTissue->AddElement(elO, 73.7723 * perCent);
brainTissue->AddElement(elNa, 0.1840 * perCent);
brainTissue->AddElement(elMg, 0.015 * perCent);
brainTissue->AddElement(elP, 0.356 * perCent);
brainTissue->AddElement(elS, 0.177 * perCent);
brainTissue->AddElement(elCl, 0.236 * perCent);
brainTissue->AddElement(elK, 0.31 * perCent);
brainTissue->AddElement(elCa, 0.009 * perCent);
brainTissue->AddElement(elFe, 0.005 * perCent);
brainTissue->AddElement(elZn, 0.001 * perCent);
// Breast
G4Material* breast = new G4Material(“Breast”, density = 0.990 * g / cm3, numberofElements = 8);
breast->AddElement(elH, 0.109);
breast->AddElement(elC, 0.506);
breast->AddElement(elN, 0.023);
breast->AddElement(elO, 0.358);
breast->AddElement(elNa, 0.001);
breast->AddElement(elP, 0.001);
breast->AddElement(elS, 0.001);
breast->AddElement(elCl, 0.001);
// Spinal Disc
G4Material* spinalDisc = new G4Material(“SpinalDisc”, 1.10 * g / cm3, numberofElements = 8);
spinalDisc->AddElement(elH, 9.60 * perCent);
spinalDisc->AddElement(elC, 9.90 * perCent);
spinalDisc->AddElement(elN, 2.20 * perCent);
spinalDisc->AddElement(elO, 74.40 * perCent);
spinalDisc->AddElement(elNa, 0.50 * perCent);
spinalDisc->AddElement(elP, 2.20 * perCent);
spinalDisc->AddElement(elS, 0.90 * perCent);
spinalDisc->AddElement(elCl, 0.30 * perCent);
// Water
G4Material* water = new G4Material(“Water”, density = 1.0 * g / cm3, numberofElements = 2);
water->AddElement(elH, 0.112);
water->AddElement(elO, 0.888);
// Muscle
G4Material* muscle = new G4Material(“Muscle”, density = 1.061 * g / cm3, numberofElements = 9);
muscle->AddElement(elH, 0.102);
muscle->AddElement(elC, 0.143);
muscle->AddElement(elN, 0.034);
muscle->AddElement(elO, 0.710);
muscle->AddElement(elNa, 0.001);
muscle->AddElement(elP, 0.002);
muscle->AddElement(elS, 0.003);
muscle->AddElement(elCl, 0.001);
muscle->AddElement(elK, 0.004);
// Liver
G4Material* liver = new G4Material(“Liver”, density = 1.071 * g / cm3, numberofElements = 9);
liver->AddElement(elH, 0.102);
liver->AddElement(elC, 0.139);
liver->AddElement(elN, 0.030);
liver->AddElement(elO, 0.716);
liver->AddElement(elNa, 0.002);
liver->AddElement(elP, 0.003);
liver->AddElement(elS, 0.003);
liver->AddElement(elCl, 0.002);
liver->AddElement(elK, 0.003);
// Tooth Dentin
G4Material* toothDentin = new G4Material(“ToothDentin”, 2.14 * g / cm3, numberofElements = 10);
toothDentin->AddElement(elH, 2.67 * perCent);
toothDentin->AddElement(elC, 12.77 * perCent);
toothDentin->AddElement(elN, 4.27 * perCent);
toothDentin->AddElement(elO, 40.40 * perCent);
toothDentin->AddElement(elNa, 0.65 * perCent);
toothDentin->AddElement(elMg, 0.59 * perCent);
toothDentin->AddElement(elP, 11.86 * perCent);
toothDentin->AddElement(elCl, 0.04 * perCent);
toothDentin->AddElement(elCa, 26.74 * perCent);
toothDentin->AddElement(elZn, 0.01 * perCent);
// Trabecular Bone
G4Material* trabecularBone =
new G4Material(“TrabecularBone”, density = 1.159 * g / cm3, numberofElements = 12);
trabecularBone->AddElement(elH, 0.085);
trabecularBone->AddElement(elC, 0.404);
trabecularBone->AddElement(elN, 0.058);
trabecularBone->AddElement(elO, 0.367);
trabecularBone->AddElement(elNa, 0.001);
trabecularBone->AddElement(elMg, 0.001);
trabecularBone->AddElement(elP, 0.034);
trabecularBone->AddElement(elS, 0.002);
trabecularBone->AddElement(elCl, 0.002);
trabecularBone->AddElement(elK, 0.001);
trabecularBone->AddElement(elCa, 0.044);
trabecularBone->AddElement(elFe, 0.001);
// Trabecular bone used in the DICOM Head
G4Material* trabecularBone_head =
new G4Material(“TrabecularBone_HEAD”, 1.18 * g / cm3, numberofElements = 12);
trabecularBone_head->AddElement(elH, 8.50 * perCent);
trabecularBone_head->AddElement(elC, 40.40 * perCent);
trabecularBone_head->AddElement(elN, 2.80 * perCent);
trabecularBone_head->AddElement(elO, 36.70 * perCent);
trabecularBone_head->AddElement(elNa, 0.10 * perCent);
trabecularBone_head->AddElement(elMg, 0.10 * perCent);
trabecularBone_head->AddElement(elP, 3.40 * perCent);
trabecularBone_head->AddElement(elS, 0.20 * perCent);
trabecularBone_head->AddElement(elCl, 0.20 * perCent);
trabecularBone_head->AddElement(elK, 0.10 * perCent);
trabecularBone_head->AddElement(elCa, 7.40 * perCent);
trabecularBone_head->AddElement(elFe, 0.10 * perCent);
// Dense Bone
G4Material* denseBone =
new G4Material(“DenseBone”, density = 1.575 * g / cm3, numberofElements = 11);
denseBone->AddElement(elH, 0.056);
denseBone->AddElement(elC, 0.235);
denseBone->AddElement(elN, 0.050);
denseBone->AddElement(elO, 0.434);
denseBone->AddElement(elNa, 0.001);
denseBone->AddElement(elMg, 0.001);
denseBone->AddElement(elP, 0.072);
denseBone->AddElement(elS, 0.003);
denseBone->AddElement(elCl, 0.001);
denseBone->AddElement(elK, 0.001);
denseBone->AddElement(elCa, 0.146);
// Cortical Bone (ICRP - NIST)
G4Material* corticalBone = new G4Material(“CorticalBone”, 1.85 * g / cm3, numberofElements = 9);
corticalBone->AddElement(elH, 4.7234 * perCent);
corticalBone->AddElement(elC, 14.4330 * perCent);
corticalBone->AddElement(elN, 4.199 * perCent);
corticalBone->AddElement(elO, 44.6096 * perCent);
corticalBone->AddElement(elMg, 0.22 * perCent);
corticalBone->AddElement(elP, 10.497 * perCent);
corticalBone->AddElement(elS, 0.315 * perCent);
corticalBone->AddElement(elCa, 20.993 * perCent);
corticalBone->AddElement(elZn, 0.01 * perCent);
// Tooth enamel
G4Material* toothEnamel = new G4Material(“ToothEnamel”, 2.89 * g / cm3, numberofElements = 10);
toothEnamel->AddElement(elH, 0.95 * perCent);
toothEnamel->AddElement(elC, 1.11 * perCent);
toothEnamel->AddElement(elN, 0.23 * perCent);
toothEnamel->AddElement(elO, 41.66 * perCent);
toothEnamel->AddElement(elNa, 0.79 * perCent);
toothEnamel->AddElement(elMg, 0.23 * perCent);
toothEnamel->AddElement(elP, 18.71 * perCent);
toothEnamel->AddElement(elCl, 0.34 * perCent);
toothEnamel->AddElement(elCa, 35.97 * perCent);
toothEnamel->AddElement(elZn, 0.02 * perCent);