Hello, dear Geant4 users.
I have written (a bit of vibe-coded) an interesting implementation for using DICOM example using browser web-based interface. Should be a good starting point for the new users which are not familiar with the C++ coding, and for the research laboratories where the end users community are larger then developer communities.
Main feature:
- Automatically fetches and installs Geant4 itself (using FetchContent option of CMake) in configure-time
- Automatically fetches and installs C++ web server (CrowCpp project, using ExternalProject of CMake, mostly because both Geant4 and Crow has similar alias for INSTALL target which is not supported directly on the end CMake side)
- Datasets (paths) might be either defined from the user-side (custom aliasing in CMake) or should be supported by default GEANT4_INSTALL_DATA=ON option (didn’t had time to check it)
- All the DICOM includes are automatically got from the sources downloaded at stage 1, none of user-defined files are used in the code
- The project deploys a two executables: orchestrator - server frontend, also creating the folder for storing the uploaded .dcm-files, and worker - the actual worker which runs the DICOM example over the user data
Output: the same output you got after running the original DICOM example.
Issues: the DCMTK option not supported (at least didn’t check as soon as didn’t want to have additional dependence here), and while original parser for non-DCMTK is not accurately bypassed, own custom header-only reader of .dcm to .g4dcm is implemented. User might upload already prepared .g4dcm from it’s own as well as .dcm that will be parser by the server to the .g4dcm.
Drawback/overhead: custom header-only http-client based on Boost::beast used for communicating between orchestrator and worker. So, Boost library is an additional dependence (also covered by FetchContent).
Geant4 version: 11.3.2 (11.4.0 release comes later then current project started, so used the old-fashioned DICOM example as a backend).
Scope of use: labs with dedicated HPC-clusters where that application might be deployed and used by many researchers for high statistics, preferably after they verify the low statistics on their workstations.
Link: github.com/mr-xak/geant4-dicom-web
License: AGPL-3.0
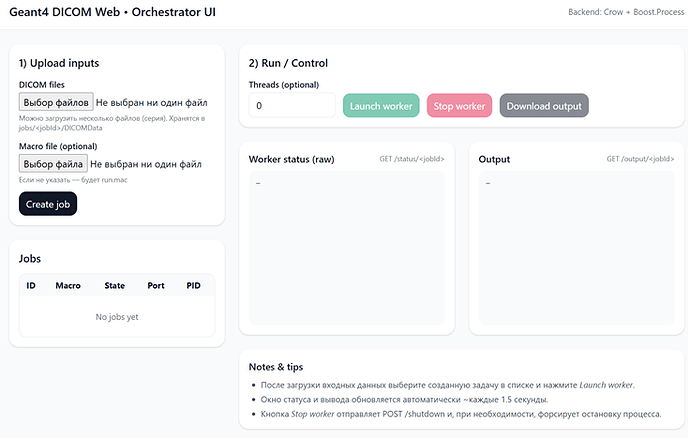
UI (vite/node.js, if NPM exists when configuring CMake, the bundle.js will be recompiled in the INSTALL target, otherwise my bundled js will be used):
Side note: conference talk (Russian) on the topic at oncoradiology.ru Friday, February 13, 2026, 16:30 MSK, Room No. 2.