Processing human_cell.C…
/Users/useruser/Geant4/geant4-v11.3.0-projects/moleculardna/./human_cell.C:314:1: warning: declaration without the ‘auto’ keyword is deprecated: function ‘__cling_Un1Qu30’ [-Wdeprecated-declarations]
SD_SSBm = sqrt(((total_SSBm2 / number) - pow(total_SSBm / number,2))/(number -1));
^~~~~~~
auto
Info in : target file: molecular-dna.root
Info in : compression setting for all output: 101
hadd Source file 1: molecular-dna_t2.root
hadd Source file 2: molecular-dna_t3.root
hadd Target path: molecular-dna.root:/
hadd Target path: molecular-dna.root:/hists
hadd Target path: molecular-dna.root:/tuples
*** Break *** segmentation violation
[/usr/lib/system/libsystem_platform.dylib] _sigtramp (no debug info)
[] (no debug info)
[] (no debug info)
[] (no debug info)
[] (no debug info)
[/opt/homebrew/Cellar/root/6.36.02/lib/root/libCling.so] cling::IncrementalExecutor::runStaticInitializersOnce(cling::Transaction&) (no debug info)
[/opt/homebrew/Cellar/root/6.36.02/lib/root/libCling.so] cling::Interpreter::executeTransaction(cling::Transaction&) (no debug info)
[/opt/homebrew/Cellar/root/6.36.02/lib/root/libCling.so] cling::IncrementalParser::commitTransaction(llvm::PointerIntPair<cling::Transaction*, 2u, cling::IncrementalParser::EParseResult, llvm::PointerLikeTypeTraitscling::Transaction*, llvm::PointerIntPairInfo<cling::Transaction*, 2u, llvm::PointerLikeTypeTraitscling::Transaction*>>&, bool) (no debug info)
[/opt/homebrew/Cellar/root/6.36.02/lib/root/libCling.so] cling::IncrementalParser::Compile(llvm::StringRef, cling::CompilationOptions const&) (no debug info)
[/opt/homebrew/Cellar/root/6.36.02/lib/root/libCling.so] cling::Interpreter::EvaluateInternal(std::__1::basic_string<char, std::__1::char_traits, std::__1::allocator> const&, cling::CompilationOptions, cling::Value*, cling::Transaction**, unsigned long) (no debug info)
[/opt/homebrew/Cellar/root/6.36.02/lib/root/libCling.so] cling::Interpreter::process(std::__1::basic_string<char, std::__1::char_traits, std::__1::allocator> const&, cling::Value*, cling::Transaction**, bool) (no debug info)
[/opt/homebrew/Cellar/root/6.36.02/lib/root/libCling.so] cling::MetaProcessor::readInputFromFile(llvm::StringRef, cling::Value*, unsigned long, bool) (no debug info)
[/opt/homebrew/Cellar/root/6.36.02/lib/root/libCling.so] TCling::ProcessLine(char const*, TInterpreter::EErrorCode*) (no debug info)
[/opt/homebrew/Cellar/root/6.36.02/lib/root/libCling.so] TCling::ProcessLineSynch(char const*, TInterpreter::EErrorCode*) (no debug info)
[/opt/homebrew/Cellar/root/6.36.02/lib/root/libCore.so] TApplication::ExecuteFile(char const*, int*, bool) (no debug info)
[/opt/homebrew/Cellar/root/6.36.02/lib/root/libRint.so] TRint::ProcessLineNr(char const*, char const*, int*) (no debug info)
[/opt/homebrew/Cellar/root/6.36.02/lib/root/libRint.so] TRint::Run(bool) (no debug info)
[/opt/homebrew/Cellar/root/6.36.02/bin/root.exe] main (no debug info)
[/usr/lib/dyld] start (no debug info)
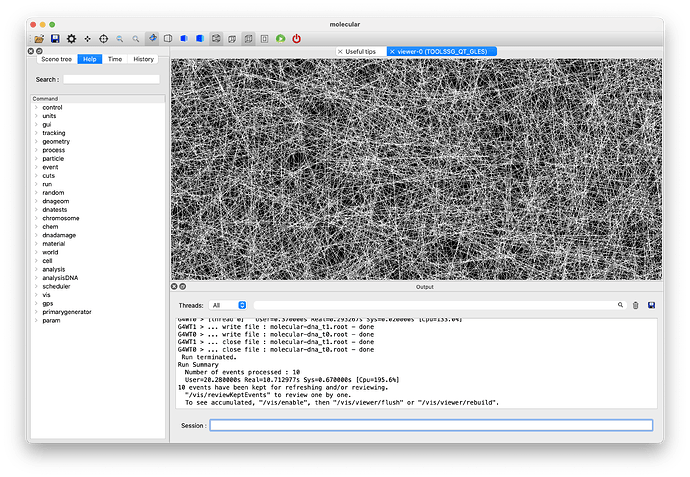
I know this is a crash (I’m a programmer) and I have no idea how I would go about debugging root, but before I investigate some more, do the previous steps and results look ok to you please?